The Mandal Lab

Research
Our Objective
The overarching goal of our lab is to unlock the potential of the gut microbiome to prevent and treat diseases. Currently, my research program is focused on three primary projects.
PROJECT 1: Investigating the Role of Gut Bacteroides in Severe Malaria
Malaria, caused by the Plasmodium parasite transmitted through female Anopheles mosquitoes, continues to pose a significant global health threat. Despite progress, the disease remains a burden, especially in sub-Saharan Africa, affecting young children which cause more than 600,000 deaths and 240 million cases annually. Our research has revealed that gut Bacteroides act in a microbial consortium to cause susceptibility to severe malaria. Bacteroides, normally beneficial gut commensals providing nutrition and supporting host immunity, are implicated in the severity of malaria. However, the mechanisms behind their involvement remain poorly understood. In this project, we aim to decipher how gut Bacteroides influence both gut and systemic immunity, exacerbating severe malaria, using a murine malaria model. Additionally, we are interested in developing a precision gut microbiota editing tool for fine-tuning the gut Bacteroides levels to promote host health.
Figure: Gut microbiota intervention strategy and possible immunological mechanism of action against severity to malaria. Agents that can modulate gut microbiota composition or deplete gut bacteria that can influence gut or systemic immunity are shown. Dotted arrow indicates the potential interactions which need further validation. Color of arrows connect gut microbiota modifying agents and their impact on respective immune cells. Role of different immune populations to inhibit or exacerbate various stage and types of malaria are connected. Although, the exact mechanism on how gut microbiota impacts severe malaria is unknown, this figure provides a plausible connection between gut microbiota and malaria severity.

PROJECT 2: Exploring the Impact of Gut Microbiome on RBC Tension, Hematopoiesis, and Severe Malaria
Our preliminary data indicate that mice susceptible to hyperparasitemia caused by Plasmodium yoelii (Py) infection exhibit increased genetic potential for the alpha hemolysin/cyclolysin transport system in their gut microbiota, compared to mice resistant to hyperparasitemia. Moreover, susceptible mice show altered expression of genes related to heme metabolism and differences in RBC surface tension compared to resistant mice. This project aims to investigate the interrelationship between alpha toxin, heme metabolism, RBC-gut axis, and severe malaria.

Atomic force microscopy (AFM) image of the red blood cell (RBC).
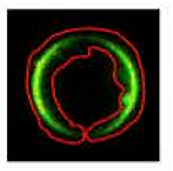
Fuorescence lifetime imaging microscopy (FLIM )microscopy of the RBC.
PROJECT 3: Decoding Species-Level Functions of Gut Commensal Bacteria Using Phage-Based Precision Perturbations
In this project, we study how individual gut commensal bacteria shape microbial community structure, metabolic function, and mucosal immunity under homeostatic conditions. Lytic bacteriophages targeting gut commensal bacteria are isolated from sewage wastewater. Using bacteriophages as highly specific ecological perturbation tools, we selectively deplete individual commensal species in vivo to reveal their causal roles in microbiome assembly, functional redundancy, and ecosystem resilience. By integrating multi-omics approaches—including metagenomics, metabolomics, transcriptomics, and immune profiling—with ecological network modeling and mouse models, we generate a systems-level understanding of cross-kingdom interactions between phages, bacteria, and the host. This work advances fundamental principles of gut microbiome organization and provides a scalable framework for precision manipulation of commensal microbes to interrogate host–microbiome symbiosis.
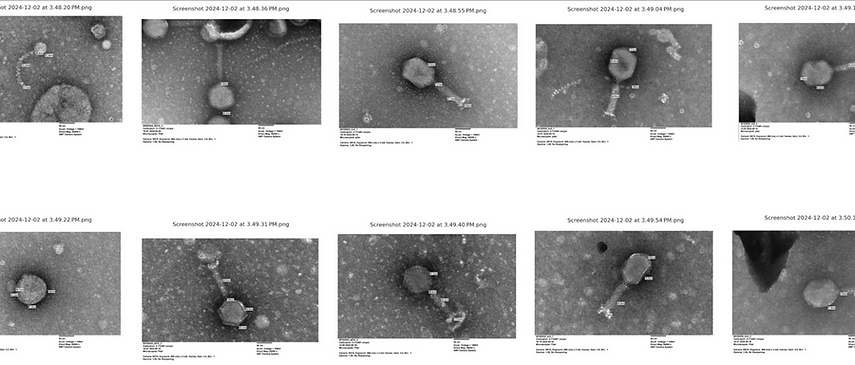

Transmission Electron Microscope (TEM) of bacteriophages against different gut commensal isolates.